About paired OAS
Sequencing of paired BCR repertoires is known to be a hard task due to the need to physically link or label natively paired variable heavy (VH) and light (VL) chains. With the advent of 10xGenomics sequencing, full-length natively paired VH/VL sequences can now be obtained, although at the expense of the B-cell throughput number in comparison to unpaired Illumina sequencing.
The Observed Antibody Space (OAS) database now provides access to annotated paired sequences from 10xGenomics studies. To date, OAS collates over 120,000 paired sequences from 7 different studies. The data is available for download or you can filter the sequences with respect to certain metadata parameters using our search form. To download the data go to the Search page.
Paired sequences in OAS can be filtered according to attributes such as species, disease, vaccination etc. The fields are non-exclusive, meaning that the user could choose a combination of fields that does not exist in our database.
Metadata
Similarly to the unpaired version of OAS, all datasets are organized into studies, that are in turn subdivided into data-units. A single data-unit is a set of sequences uniquely identified by its metadata. The range of meta-parameters are:
- Age Information on age of the human B-cell donors.
- Disease Indicates whether the donor was sick at the time of B-cell extraction.
- Vaccine Indicates whether the B-cell donor was purposely immunised prior to B-cell extraction.
- BType Indicates whether B-cells were experimentally sorted prior to sequencing.
- Species Organism of the B-cell donor.
- Author First author and the publication year.
- Unique sequences Number of sequences that passed quality filtering steps.
- BSource Which organ/tissue the B-cells were extracted from.
- Subject Indicates whether the B-cells can be tracked back to a particular individual.
- Longitudinal If the study is conducted over a period of time, indicates the particular timepoint when B-cells were sourced.
Data Format
A series of .csv.gz files will then be downloaded to your current directory. Each .csv.gz file contains as the first line the metadata for the data-unit and the following lines each sequence and its annotations.
The contents of each data-unit file can look as below:
| sequence_id_heavy | sequence_heavy | locus _heavy |
stop_codon _heavy |
vj_in_frame _heavy |
productive _heavy |
rev_comp _heavy |
v_call _heavy |
d_call _heavy |
j_call _heavy |
sequence_alignment _heavy |
germline_alignment _heavy |
sequence_alignment_aa _heavy |
germline_alignment_aa _heavy |
v_alignment _start_heavy |
v_alignment _end_heavy |
d_alignment _start_heavy |
d_alignment _end_heavy |
j_alignment _start_heavy |
j_alignment _end_heavy |
v_sequence_alignment _heavy |
v_sequence_alignment_aa _heavy |
v_germline_alignment _heavy |
v_germline_alignment_aa _heavy |
d_sequence_alignment _heavy |
d_sequence_alignment _aa_heavy |
d_germline_alignment _heavy |
d_germline_alignment _aa_heavy |
j_sequence_alignment _heavy |
j_sequence_alignment_aa _heavy |
j_germline_alignment _heavy |
j_germline_alignment_aa _heavy |
fwr1 _heavy |
fwr1_aa _heavy |
cdr1 _heavy |
cdr1_aa _heavy |
fwr2 _heavy |
fwr2_aa _heavy |
cdr2 _heavy |
cdr2_aa _heavy |
fwr3 _heavy |
fwr3_aa _heavy |
cdr3 _heavy |
cdr3_aa _heavy |
junction _heavy |
junction _length_heavy |
junction_aa _heavy |
junction_aa _length_heavy |
v_score _heavy |
d_score _heavy |
j_score _heavy |
v_cigar _heavy |
d_cigar _heavy |
j_cigar _heavy |
v_support _heavy |
d_support _heavy |
j_support _heavy |
v_identity _heavy |
d_identity _heavy |
j_identity _heavy |
v_sequence _start_heavy |
v_sequence _end_heavy |
v_germline _start_heavy |
v_germline _end_heavy |
d_sequence _start_heavy |
d_sequence _end_heavy |
d_germline _start_heavy |
d_germline _end_heavy |
j_sequence _start_heavy |
j_sequence _end_heavy |
j_germline _start_heavy |
j_germline _end_heavy |
fwr1_start _heavy |
fwr1_end _heavy |
cdr1_start _heavy |
cdr1_end _heavy |
fwr2_start _heavy |
fwr2_end _heavy |
cdr2_start _heavy |
cdr2_end _heavy |
fwr3_start _heavy |
fwr3_end _heavy |
cdr3_start _heavy |
cdr3_end _heavy |
np1 _heavy |
np1_length _heavy |
np2 _heavy |
np2_length _heavy |
sequence_id _light |
sequence _light |
locus _light |
stop_codon _light |
vj_in_frame _light |
productive _light |
rev_comp _light |
v_call _light |
d_call _light |
j_call _light |
sequence_alignment _light |
germline_alignment _light |
sequence_alignment_aa _light |
germline_alignment_aa _light |
v_alignment _start_light |
v_alignment _end_light |
d_alignment _start_light |
d_alignment _end_light |
j_alignment _start_light |
j_alignment _end_light |
v_sequence_alignment _light |
v_sequence_alignment_aa _light |
v_germline_alignment _light |
v_germline_alignment_aa _light |
d_sequence_ alignment_light |
d_sequence_ alignment_aa_light |
d_germline_ alignment_light |
d_germline_ alignment_aa_light |
j_sequence_ alignment_light |
j_sequence_alignment _aa_light |
j_germline_alignment _light |
j_germline_alignment _aa_light |
fwr1 _light |
fwr1_aa _light |
cdr1 _light |
cdr1_aa _light |
fwr2 _light |
fwr2_aa _light |
cdr2 _light |
cdr2_aa _light |
fwr3 _light |
fwr3_aa_ light |
cdr3 _light |
cdr3_aa _light |
junction _light |
junction_length _light |
junction_aa _light |
junction_aa_ length_light |
v_score _light |
d_score _light |
j_score _light |
v_cigar _light |
d_cigar _light |
j_cigar _light |
v_support _light |
d_support _light |
j_support_light | v_identity _light |
d_identity _light |
j_identity _light |
v_sequence _start_light |
v_sequence _end_light |
v_germline _start_light |
v_germline _end_light |
d_sequence _start_light |
d_sequence _end_light |
d_germline _start_light |
d_germline _end_light |
j_sequence _start_light |
j_sequence _end_light |
j_germline _start_light |
j_germline _end_light |
fwr1_start _light |
fwr1_end _light |
cdr1_start _light |
cdr1_end _light |
fwr2_start _light |
fwr2_end _light |
cdr2_start _light |
cdr2_end _light |
fwr3_start _light |
fwr3_end _light |
cdr3_start _light |
cdr3_end _light |
np1 _light |
np1_length _light |
np2 _light |
np2_length _light |
ANARCI_numbering_light | ANARCI_numbering_heavy | ANARCI_status _light |
ANARCI_status _heavy |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AAACCTGAGGGATACC-1 _contig_1 |
ACGGAGGTTTCT... | IGH | F | T | T | F | IGHV9-3*01 | IGHD2-4*01 | IGHJ3*01 | CAGATCCAGTTGG... | CAGATCCAGTTG... | QIQLVQSGPELKKPG... | QIQLVQSGPELKKPG... | 1.0 | 290.0 | 295.0 | 303.0 | 306.0 | 349.0 | CAGATCCAGTTGGT... | QIQLVQSGPELKKPGETV... | CAGATCCAGTTGGT... | QIQLVQSGPELKKPGE... | GATTACGAC | DYD | GATTACGAC | DYD | GTTTGCTTACTGGG... | FAYWGQGTLVTVSA | GTTTGCTTACTGGG... | FAYWGQGTLVTVSA | CAGATCCAG... | QIQLVQ... | GGGTATAC... | GYTFTTYG | ATGAGCT... | MSWVKQ... | ATAAAC... | INTYSGVP | ACATATG... | TYVDDFK... | GCCCCC... | APDYDEFAY | TGTGCCC... | 33.0 | CAPDYDE... | 11.0 | 450.572 | 17.992 | 85.286 | 309S290M150S4N | 603S8N9M137S | 614S4N44M91S | 2.040000e-128 | 0.839300 | 1.019000e-20 | 99.655 | 100.0 | 100.0 | 310.0 | 599.0 | 1.0 | 290.0 | 604.0 | 612.0 | 9.0 | 17.0 | 615.0 | 658.0 | 5.0 | 48.0 | 310.0 | 384.0 | 385.0 | 408.0 | 409.0 | 459.0 | 460.0 | 483.0 | 484.0 | 597.0 | 598.0 | 624.0 | CCCC | 4.0 | GA | 2.0 | AAACCTGAGGGATACC-1_contig_2 | TTGGGAGGG... | IGK | F | T | T | F | IGKV1-117*01 | NaN | IGKJ1*01 | GATGTTTTGATGAC... | GATGTTTTGATG... | DVLMTQTPLSLPVSLGD... | DVLMTQTPLSLPVSL... | 1.0 | 299.0 | NaN | NaN | 300.0 | 337.0 | GATGTTTTGATGACCC... | DVLMTQTPLSLPVSLGDQ... | GATGTTTTGATGAC... | DVLMTQTPLSLPVSLGDQ... | NaN | NaN | NaN | NaN | GTGGACGTTC... | WTFGGGTKLEIK | GTGGACGTTCGGT... | WTFGGGTKLEIK | GATGTTT... | DVLMTQT... | CAGAGCA... | QSIVHSNGNTY | TTAGAATG... | LEWYLQ... | AAAGTTTCC | KVS | AACCGATTT... | NRFSGVPDRFSGS... | TTTCAAGG... | FQGSHLPWT | TGCTTTCA... | 33.0 | CFQGSHLPWTF | 11.0 | 464.595 | NaN | 73.749 | 140S299M121S3N | NaN | 439S38M83S | 9.090000e-133 | NaN | 2.324000e-17 | 99.666 | NaN | 100.000 | 141.0 | 439.0 | 1.0 | 299.0 | NaN | NaN | NaN | NaN | 440.0 | 477.0 | 1.0 | 38.0 | 141.0 | 218.0 | 219.0 | 251.0 | 252.0 | 302.0 | 303.0 | 311.0 | 312.0 | 419.0 | 420.0 | 446.0 | NaN | 0.0 | NaN | NaN | {'fwl1': {'1': 'D', '2': 'V', '3': 'L', '4': 'M', ...}, 'cdrl1': {'27': 'Q', '28': 'S', '29': 'I', '30': 'V', ...} ...'cdrl3': {'105': 'F', '106': 'Q', '107': 'G' ...}, |
{'fwh1': {'1': 'Q', '2': 'I', '3': 'Q' ...}, 'cdrh1': {'27': 'G', '28': 'Y',...} ...'cdrh3': {'105': 'A', '106': 'P', ...}, |
||||| | ||||Shorter than IMGT defined: fw4| |
| AAACCTGCACAGATTC-1 _contig_2 |
TGAAAACAACCT... | IGH | F | T | T | F | IGHV1-81*01 | IGHD1-1*01 | IGHJ1*03 | CAGGTTCAGCTGC... | CAGGTTCAGCT... | QVQLQQSGAELARP... | QVQLQQSGAELARP... | 1.0 | 294.0 | 305.0 | 318.0 | 322.0 | 373.0 | CAGGTTCAGCTGC... | QVQLQQSGAELARPGAS... | CAGGTTCAGCTGCA... | QVQLQQSGAELARPGA... | TTTATTACTAC... | YYYG | TTTATTACTACGGT | YYYG | TACTGGTACTTCGAT... | YWYFDVWGTGTTVTVSS | TACTGGTACTTCGA... | YWYFDVWGTGTTVTVSS | CAGGTTCAG... | QVQLQ... | GGCTACAC... | GYTFTSYG | ATAAGCT... | ISWVKQR... | ATTTATC... | IYLRSGNT | TACTACA... | YYNEKFK... | GCAAGA... | ARWERFY... | TGTGCAA... | 57.0 | CARWERFYY... | 19.0 | 456.805 | 27.605 | 100.667 | 115S294M170S | 419S14M146S9N | 436S1N52M91S | 2.073000e-130 | 0.000825 | 1.897000e-25 | 99.660 | 100.0 | 100.0 | 116.0 | 409.0 | 1.0 | 294.0 | 420.0 | 433.0 | 1.0 | 14.0 | 437.0 | 488.0 | 2.0 | 53.0 | 116.0 | 190.0 | 191.0 | 214.0 | 215.0 | 265.0 | 266.0 | 289.0 | 290.0 | 403.0 | 404.0 | 454.0 | TGGGAG... | 10.0 | TCT | 3.0 | AAACCTGCACAGATTC- 1_contig_1 |
GAAATACATC... | IGK | F | T | T | F | IGKV6-23*01 | NaN | IGKJ2*01 | GACATTGTGATGAC... | GACATTGTGATG... | DIVMTQSHKFMSTSVGD... | DIVMTQSHKFMSTSV... | 1.0 | 283.0 | NaN | NaN | 284.0 | 319.0 | GACATTGTGATGACCC... | DIVMTQSHKFMSTSVGDR... | GACATTGTGATGAC... | DIVMTQSHKFMSTSVGDR... | NaN | NaN | NaN | NaN | ACACATTCGG... | TFGGGTKLEIK | ACACGTTCGGAGG... | TFGGGTKLEIK | GACATTG... | DIVMTQS... | CAGGATG... | QDVGTA | GTAGCCT... | VAWYQQ... | TGGGCATCC | WAS | ACCCGGCA... | TRHTGVPDRFTGS... | CAGCAATAT... | QQYSNYHT | TGTCAGCA... | 30.0 | CQQYSNYHTF | 10.0 | 433.433 | NaN | 64.136 | 90S283M119S4N | NaN | 373S3N36M83S | 1.905000e-123 | NaN | 1.595000e-14 | 98.940 | NaN | 97.222 | 91.0 | 373.0 | 1.0 | 283.0 | NaN | NaN | NaN | NaN | 374.0 | 409.0 | 4.0 | 39.0 | 91.0 | 168.0 | 169.0 | 186.0 | 187.0 | 237.0 | 238.0 | 246.0 | 247.0 | 354.0 | 355.0 | 378.0 | NaN | 0.0 | NaN | NaN | {'fwl1': {'1': 'D', '2': 'I', '3': 'V', '4': 'M', '5': }, 'cdrl1': {'27': 'Q', '28': 'D', '29': 'V', '36': 'G', ...} ...'cdrl3': {'105': 'Q', '106': 'Q', '107': 'Y' ...}, |
{'fwh1': {'1 ': 'E', '2 ': 'V', '3 ': 'Q', '4': ... }, 'cdrh1': {'27': 'G', '28': 'Y', '29': ...} ... cdrh3': {'105': 'A', '106': 'R', '107': ...}, |
'|||||' | '||||Shorter than IMGT defined: fw4|' |
Linking heavy and light chains
In the ideal case scenario, 10xGenomics sequencing would yield 1-to-1 heavy/light chain pairings for each interrogated B-cell. However, in many cases more than one heavy and/or light chain sequences harbour identical 10xGenomics cell barcodes. Linking such heavy and light chain sequences can lead to combinatorial inflation of the real sequence number and incorrect estimation of the repertoire diversity (Figure 2).
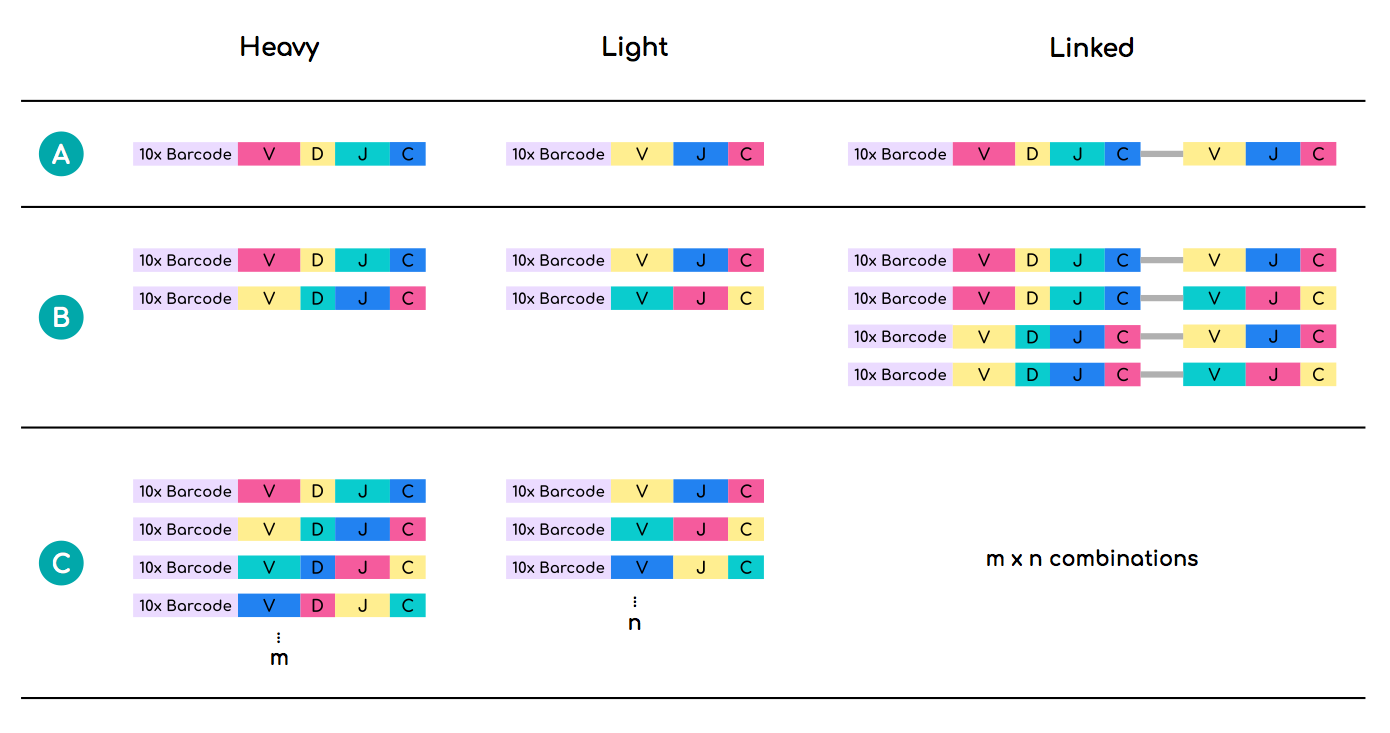
One solution is to filter out sequences whose 10xGenomics barcodes are shared between more than one unique heavy and light V(D)J recombination events. This step has already been performed for each data unit in OAS. However, a recent study showed that individual B-cells can produce more than two functional V(D)J recombinations (Shi et al., 2019. Cell Discovery).
Contact
If you would like to contact us or to submit your study to OAS, please drop an email to oas_opig@stats.ox.ac.uk.
